Gene duplication
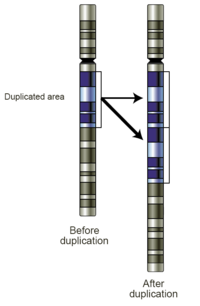
Gene duplication (or gene amplification) is any duplication of a region of DNA that contains a gene.
Gene duplication is important in supplying raw genetic material – new genes – to biological evolution. This has been recognized since the 1930s. Recent genomic sequence data show that duplicated genes are common in all organisms surveyed.[1]
Origin[change | change source]
In classical genetics it may occur as an error in crossing-over during meiosis, or the duplication of an entire chromosome.[1]
In non-classical genetics it may happen by a retrotransposition event.[1]
Implications[change | change source]
The second copy of the gene is often relatively free from selective pressure. Mutations of it have no deleterious effects to its host organism. Thus it accumulates mutations faster than a functional single-copy gene, over generations.
A duplication is the opposite of a deletion. Duplications arise from unequal crossing-over that occurs during meiosis between misaligned homologous chromosomes. The product of this recombination are a duplication at the site of the exchange and a reciprocal deletion.[2]
Evolutionary importance[change | change source]
Gene duplication is believed to play a major role in evolution; this idea was first suggested 100 years ago.[3] Susumu Ohno was one of the most famous developers of this theory in his classic book Evolution by gene duplication (1970).[4] Ohno argued that gene duplication is the most important evolutionary force since the emergence of the last universal common ancestor (LUCA).[5]
Major genome duplication events are not uncommon. It is believed that the entire yeast genome underwent duplication about 100 million years ago.[6] Plants are the most prolific genome duplicators. For example, wheat is hexaploid (a kind of polyploid), meaning that it has six copies of its genome.
Normally, a big change in gene function is resisted because the original function is needed, but after a duplication one gene carries on the original function. Therefore, a change of function in the second copy is possible without loss of fitness.[7][8][9] This freedom from consequences allows for more mutations to be 'carried' in the population. Some of these might increase the fitness of the organism or code for a new function.
The two genes that exist after a gene duplication event are called paralogs and usually code for proteins with a similar function and/or structure. By contrast, orthologous genes are ones which code for proteins with similar functions but exist in different species, and are created from species-splitting.
Related pages[change | change source]
References[change | change source]
- ↑ 1.0 1.1 1.2 Zhang J (2003). "Evolution by gene duplication: an update". Trends in Ecology & Evolution. 18 (6): 292–8. doi:10.1016/S0169-5347(03)00033-8.
- ↑ "Definition of Gene duplication". MedicineNet. Archived from the original on 2014-03-06. Retrieved 2019-02-14.
- ↑ Taylor J.S. Raes J. (2004). "Duplication and divergence: the evolution of new genes and old ideas". Annu. Rev. Genet. 38: 615–43. doi:10.1146/annurev.genet.38.072902.092831. PMID 15568988.
- ↑ Ohno, S. (1970). Evolution by gene duplication. Springer-Verlag. ISBN 0-04-575015-7.
- ↑ Ohno, S. (1967). Sex chromosomes and sex-linked genes. Springer-Verlag. ISBN 91-554-5776-2.
- ↑ Kellis M, Birren BW, Lander ES (April 2004). "Proof and evolutionary analysis of ancient genome duplication in the yeast Saccharomyces cerevisiae". Nature. 428 (6983): 617–24. Bibcode:2004Natur.428..617K. doi:10.1038/nature02424. PMID 15004568. S2CID 4422074.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ↑ De Smet R. & Van de Peer Y. 2012. Redundancy and rewiring of genetic networks following genome-wide duplication events. Current opinion in plant biology. Feb, 1-9.
- ↑ Ruby J.G. et al 2007. Evolution, biogenesis, expression, and target predictions of a substantially expanded set of Drosophila microRNAs. Genome research 17 (12) 1850-1864.
- ↑ D.H. Graur and W.-H.C. Li 2000. Fundamentals of Molecular Evolution. 2nd ed, Sinauer.
