CRISPR

1. cas genes,
2. a leader sequence, and
3. A repeat-spacer array.
The arrangement of the three components is not always as shown

CRISPR is a term in DNA research. It stands for clustered regularly-interspaced short palindromic repeats. These are part of the genetic code in prokaryotes: most bacteria and archaea have it. It is their defence against attack by viruses.[1] Its structure and function was discovered in the 21st century.[2][3][4]
CRISPR has a lot of short repeated sequences. These sequences are part of an adaptive immune system for prokaryotes. It allows them to remember and counter the bacteriophages which prey on them. They work as a kind of acquired immunity for bacteria.[5][6]
They can modify the genes of almost any organism. They are used by researchers as a tool to cut and insert genes in genetic modification (GM).[7] Work is under way to find how they can be used to attack virus diseases in humans (gene therapy).[8][9]
How it works
[change | change source]Each repetition is followed by short segments of "spacer DNA". These come from previous exposures to a bacterial virus or plasmid.[8] CRISPR spacers recognize and cut up the foreign genetic elements in a manner like RNA interference in eukaryotic organisms.
In effect, the spacers are fragments of DNA from viruses that have previously tried to attack the cell line. The foreign source of the spacers was a sign to researchers that the CRISPR/cas system could have a role in adaptive immunity in bacteria.[10]
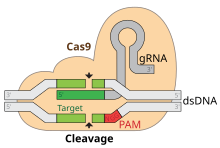
The actual cutting is done by a nuclease called Cas9. Cas9 has two active cutting sites, one for each strand of the DNA's double helix. Cas9 does this by unwinding foreign DNA and checking whether it is complementary to the 20 basepair spacer region of the guide RNA (the spacer region RNA). If it is, the foreign DNA gets chopped up.[7][11]
Applications
[change | change source]CRISPR is an anti-viral defensive system which originated in bacteria and archaea. Gene editing is a human exploitation of CRISPR.
The technology has been used to switch off genes in human cell lines and cells, to study Candida albicans, to modify yeasts used to make biofuel and to genetically modify crop strains.[12]
The CRISPR-Cas9 system cuts DNA, but can do more. It can turn gene expression on and off, and can be used to fluorescently tag particular sequences.[13][14]
Nobel award
[change | change source]The understanding and development of the CRISPR-Cas9 genome editing technique won the Nobel Prize in Chemistry for 2020. The prize was awarded jointly to Emmanuelle Charpentier and Jennifer Doudna.[15][16]
Anti-CRISPR
[change | change source]Anti-CRISPR is a group of proteins in some phages. It inhibits the normal activity of CRISPR-Cas, the immune system of many bacteria. Phages with anti-CRSPR avoid having their genomes destroyed by the prokaryotic cells that they infect.[17]
References
[change | change source]- ↑ Grissa I; Vergnaud G. & Pourcel C. 2007. The CRISPRdb database and tools to display CRISPRs and to generate dictionaries of spacers and repeats. BMC Bioinformatics 8: 172. [1]
- ↑ Gesner E.M; Schellenberg M.J; Garside E.L; George M.M. & Macmillan A.M. 2011. Recognition and maturation of effector RNAs in a CRISPR interference pathway. Nature Structural & Molecular Biology. 18 (6): 688–692.
- ↑ See Nobel Prize in Chemistry to Emmanuelle Charpentier and Jennifer Doudna, 2020.
- ↑ Wiedenheft B; Sternberg S.H; Doudna J.A. 2012. RNA-guided genetic silencing systems in bacteria and archaea. Nature. 482 (7385): 331–338.[2]
- ↑ Lecture by Jennifer Doudna [3]
- ↑ Mojica F.J.M. et al 2000. Biological significance of regularly spaced repeats in the genomes of Archaea, Bacteria and mitochondria. Molecular Microbiology 36: 244–246.
- ↑ 7.0 7.1 Carey, Nessa 2019. Hacking the code of life: how gene editing will rewrite our futures. London: Icon Books. ISBN 978-178578-625-9
- ↑ 8.0 8.1 Marraffini L.A. & Sontheimer E.J. 2010. CRISPR interference: RNA-directed adaptive immunity in bacteria and archaea. Nature Reviews Genetics 11 (3): 181–190. [4]
- ↑ Hille F. & Charpentier E. 2016. CRISPR-Cas: biology, mechanisms and relevance. Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences. 371 (1707): 20150496. [5]
- ↑ Morange, M (2015). "What history tells us XXXVII. CRISPR-Cas: The discovery of an immune system in prokaryotes". J. Biosci. 40 (2): 221–223. doi:10.1007/s12038-015-9532-6. PMID 25963251. S2CID 14048176.
- ↑ Re-writing the Code of Life: CRISPR systems and applications of gene editing. Jennifer Doudna at the Royal Society on YouTube. [6]
- ↑ Ledford H (2015). "CRISPR, the disruptor". Nature. 522 (7554): 20–4. Bibcode:2015Natur.522...20L. doi:10.1038/522020a. PMID 26040877. S2CID 4456595.
- ↑ CRISPR: Gene editing and beyond/ [7]
- ↑ Doudna, Jennifer & Sternberg, Samuel 2017. A crack in creation: the new power to control evolution. London: Bodley Head. ISBN 978-1-847-92382-0
- ↑ "Press release: The Nobel Prize in Chemistry 2020". Nobel Foundation. Retrieved 7 October 2020.
- ↑ Wu KJ, Peltier E (7 October 2020). "Nobel Prize in Chemistry Awarded to 2 Scientists for Work on Genome Editing – Emmanuelle Charpentier and Jennifer A. Doudna developed the Crispr tool, which can alter the DNA of animals, plants and microorganisms with high precision". The New York Times. Retrieved 7 October 2020.
- ↑ Stanley SY, Borges AL, Chen KH, Swaney DL, Krogan NJ, Bondy-Denomy J, Davidson AR 2019. Anti-CRISPR-associated proteins are crucial repressors of Anti-CRISPR transcription. Cell 178 (6): 1452–1464.e13. doi:10.1016/j.cell.2019.07.046. PMC 6754177. PMID 31474367
